This course covers the computational methods for the analysis of genome wide association studies and next generation
sequencing technologies. The goal of the course is to provide training in interdisciplinary computational analysis,
to learn how to identify computational problems/solutions, and to make contributions in other fields without an
extensive background.
- The Computational Problem
-
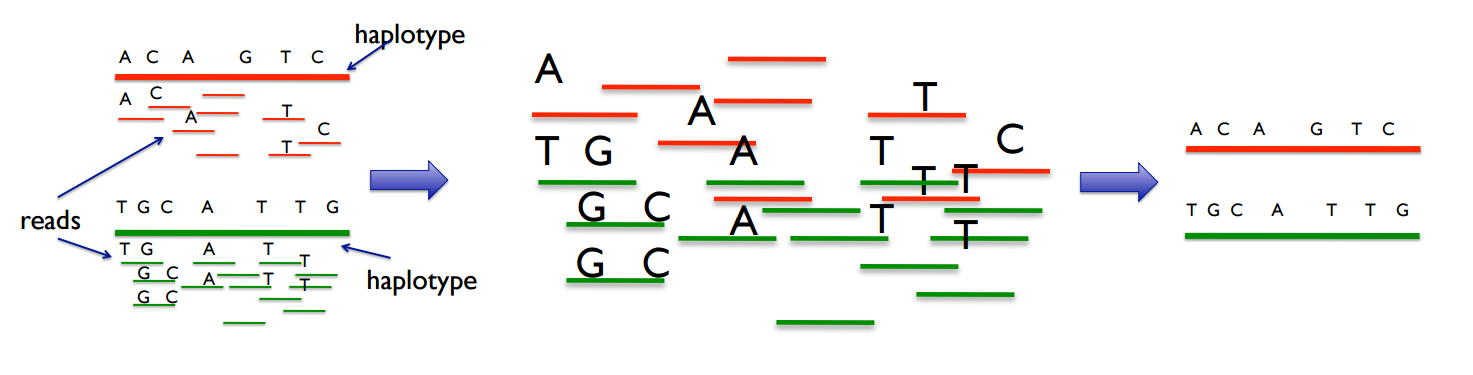
Humans have 2 haplotypes; while sequencers produce reads that only contain regional info from one haplotype.
-
We don't know which read is from which haplotype.
-
But by aligning and using info from multiple reads, we can reassemble the haplotypes. (See header diagram).
-
This is called the Haplotype Assembly Problem.
- Input Format
-
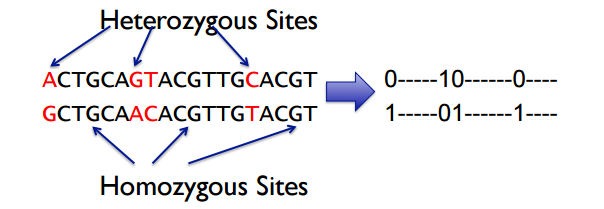
A pair of haplotypes are represented as a pair of complementary binary strings.
Homozygous sites are not represented, and heterozygous sites are assigned 0/1 pairings.
-
Reads are thus also represented as binary strings.
-
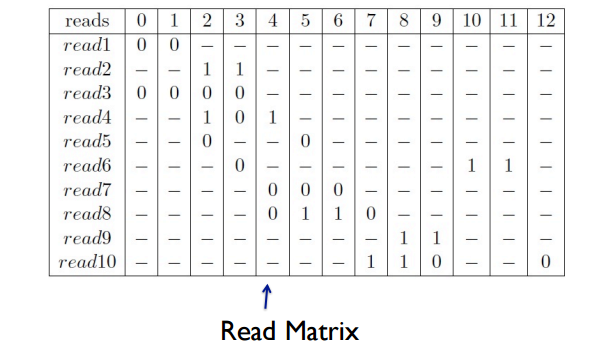
Multiple reads are then a set of binary strings, which can be represented as a binary read matrix.
-
This read matrix can then be analyzed to determine haplotype pairs/sets.
- Output Format
-
Output format will be the pair of complementary binary strings representing the haplotypes.
Below is a possible solution to the above read matrix.

- Algorithm Benchmarks
-
In order to benchmark different algorithms, we can analyze each one's space and time complexities for solving the computational problem.
-
Time complexity is based on the number of operations that the algorithm will end up performing.
-
Space complexity is based on the temporary storage required by the algorithm (this doesn't include the input read matrix).
-
Let the number of SNPs/sites (length of each sequence) be N.
-
Let the number of reads be R.
-
Let the number of haplotypes be H.
- Baseline Solution Algorithms
-
Trivial Guess-and-Check Algorithm (Easy Difficulty)
-
Assume all reads are correct (no errors) and only two haplotypes (H = 2). Iterate through all
possible haplotype combinations, and verify with all reads to
check if it's a solution.
-
Time Complexity: < R * 2N = O(2N)
-
Space Complexity: = 2 * N = O(N)
-
Problem(s):
-
Extremely inefficient. Exponential time complexity.
-
If there are any error reads, it might not solve.
-
Doesn't account for other numbers of haplotypes.
-
Trivial Try-All-Choose-Best Algorithm (Medium Difficulty)
-
Allow for read errors and assume only two haplotypes (H = 2). Iterate through all possible haplotype
combinations, and count how many read errors each one would have.
Store each combination's read errors in an array. Choose the
combination that would produce the least number of errors.
-
Time Complexity: = R * 2N = O(2N)
-
Space Complexity: = O(2N)
-
Problem(s):
-
Extremely inefficient. Exponential time & space complexities.
-
Doesn't account for other numbers of haplotypes.
- Improved Solution Algorithms
-
Greedy Algorithm (Easy Difficulty)
-
Assume all reads are correct (no errors) and only two haplotypes (H = 2). For each SNP of the haplotype sequence, copy the read until end of read is found '-', save that index.
Move to next read with non-hyphen at that index. Check which haplotype this read belongs to by examining a previous SNP.
Continue copying the read until end, and save index again. Repeat until end of the sequence.
-
Time Complexity: = R + N = O(N), assuming N >> R
-
Space Complexity: = 2 * N = O(N)
-
Problem(s):
-
If there are any error reads, it might not solve.
-
Doesn't account for other numbers of haplotypes.
-
Greedy Algorithm (Medium Difficulty)
-
Allow for read errors and assume only two haplotypes (H = 2). Examine coinciding SNP regions for a local
solution that minimizes local errors. Repeat until entire sequence is generated.
-
Time Complexity: < R + N = O(N), assuming N >> R
-
Space Complexity: < (2 * N) + R = O(N), assuming N >> R
-
Problem(s):
-
Doesn't account for other numbers of haplotypes.
- Weekly Updates
-
Week 7
-
Progress: Project Decision and Website Wiki Setup
-
Next Week: Description of Computational Problem
-
Grade for this Week: A+
-
Problems that came up: None
-
Problems solved: Getting a webpage wiki framework setup
-
Week 8
-
Progress: Description of Computational Problem
-
Next Week: Definition of Inputs & Output Benchmarks
-
Grade for this Week: A+
-
Problems that came up: None
-
Problems solved: Outlining what Haplotype Assembly entails
-
Week 9
-
Progress: Definition of Inputs & Output Benchmarks
-
Next Week: Trivial/Improved Algorithm (Easy)
-
Grade for this Week: A+
-
Problems that came up: None
-
Problems solved: Input & Output format description
-
Week 10
-
Progress: Trivial/Improved Algorithms (Easy)
-
Next Week: Attempt to Allow for Read Errors (Medium)
-
Grade for this Week: A+
-
Problems that came up: None
-
Problems solved: Outlined and benchmarked algorithm solutions
-
Week 11
-
Progress: Attempt to Allow for Read Errors (Medium)
-
Grade for this Week: A for Effort
-
Problems that came up: Medium was harder than originally thought. Outlined idea, but no implementation.
-
Problems solved: Presentation, Project Packaging, and Documentation